The neuroinformatics of neural connectivity
- Advances in the automatic analysis of multi-dimensional data
- Ontologies for neuroscience: applications and advances
- How should a neuron be modeled? Biophysical detail vs. abstraction
- High performance computing and grid infrastructure for neuroinformatics applications
- The neuroinformatics of neural connectivity
Workshop 5
Title: The neuroinformatics of neural connectivity
Chair: David Willshaw, University of Edinburgh, UK
Co-chair: Kevan Martin, University of Zurich, Switzerland
The brain is perhaps unique amongst the organs of the body in that its major components, the nerve cells, are highly interconnected. Typically each neuron has many thousands of synapses and the distribution of synapses on branching dendritic trees of individual neurons may underly complex local processing within neurons. These intricate patterns of connectivity enables sophisticated processing within single cells, groups of cells and brain areas, so contributing to the remarkable computational capabilities of the brain, unmatched by any manmade machine.
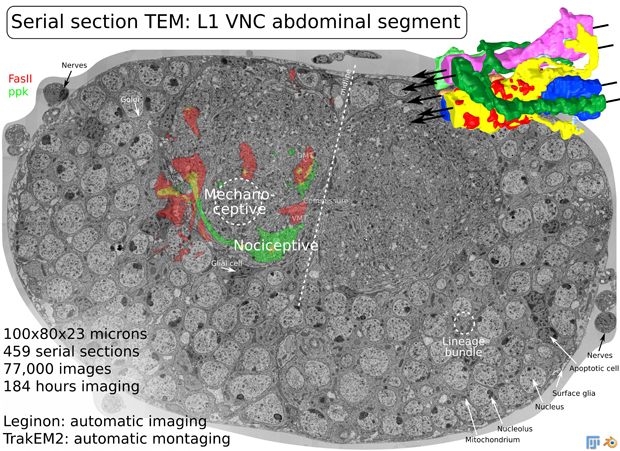
The complexity of brain connectivity makes it a particular challenge for understanding the development and functioning of the brain in normal and in damaged or pathological states. In this workshop we examine neuroinformatics approaches to an analysis of brain connectivity at three different levels: the ultrastructural level, the cellular level and the brain imaging level.
Speakers:
-
Albert Cardona, University of Zurich/ETH Zurich, Switzerland
-
Giorgio Ascoli, George Mason University, USA
-
Rolf Kötter, Radboud University Nijmegen, Netherlands